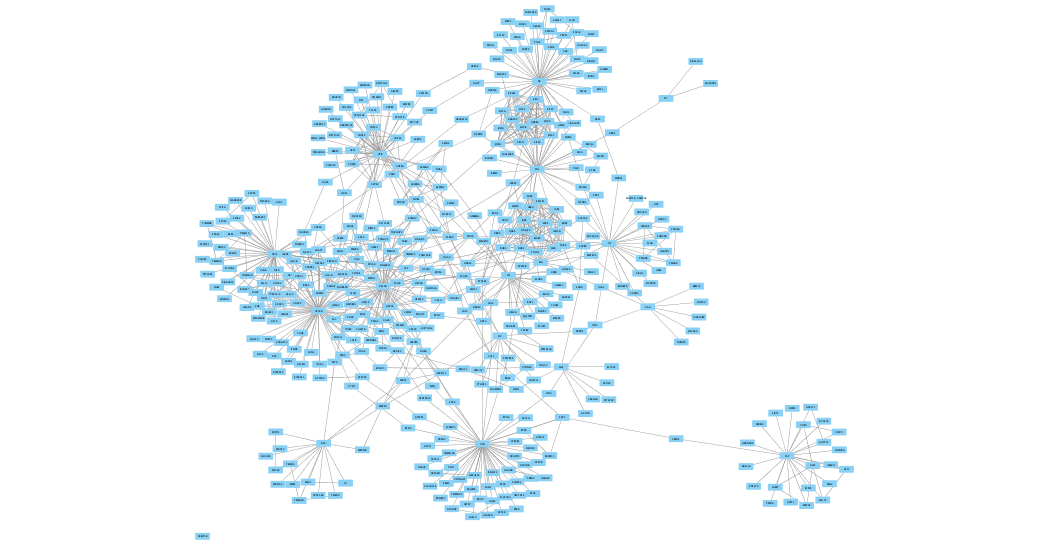
Next, the Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analyses were dissected using R language. In this study, the potential targets of compounds identified in SYP were predicted by Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP), and a “herb-compound-target” network was constructed via Cytoscape. The renal protective mechanisms of Shenyuan particle (SYP) in the treatment of diabetic kidney disease (DKD) were investigated, focusing on the main targets and pathways. As you can see in VizMapper, there are a multitude of options for coloring and otherwise changing your network.Here, we can see that the shared OTUs are colored yellow, and the not_shared OTUs are colored pink. Now you can click on Node Color and color each individually. With this information, make a new column in your real_node_table.txt where you label each OTU based on it being shared or not_shared, and each sample based on being a Control or Fasting sample. You can now use the COUNTIF function in Excel to find if each OTU is present in either the Control group only, the Fasting group only, or in both. In column C put your entire list of OTUs taken from the otu_table.txt. Make a new excel sheet that contains where column A represents the OTUs seen in the Control group, column B represents the samples in Fasting group.Open up your real_edge_table.txt in Excel.Copy out all the OTU Ids seen in your study.Convert your biom table to a classic otu table using the biom convert command. 
For example, how can we color the otus that are shared between the Control and Fasting groups a different color than the otus that belong to just one treatment group? To do this, we can:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
